Introduction
- It is called as soluble RNA because of its small size made up of 75-95 nucleotides and sedimentation coefficient of mature eukaryotic t-RNA is 3.8 S.
- There are about 60 different specific t-RNAs in bacterial and 100-110 in mammalian cells.
- t-RNA is synthesized on a DNA template and is an exception to other cellular RNAs in that a part of it as ribonucleotide sequence (-CCA) is added after it comes off DNA template.
- Each amino-acid is carried by specific t-RNA.
- Since 20 amino-acids are coded to form proteins, there must be at least 20 types of t-RNA.
- However, it has been found that there are at least 2 types for each amino-acid signifying their higher number.
- t-RNAs are the adaptors between codons and amino-acids they specify.
- Most t-RNA recognizes more than one codon but such codons should specify a particular amino acid.
- The 3’ terminal has –CCA sequence where amino acid is attached.
- There are various unusual bases in t-RNA which are pseudouridine (ΨU), dihydrouridine, Hypoxanthine, Methylguanine, Dimethyl guanosine, Inosine, Methyl inosine, Ribothymidine, etc.
- Thee bases improve the t-RNA function. E.g., Hypoxanthine play an important role in codon recognition.
Clover leaf: secondary structure
- The secondary structure of t-RNA shows various parts. They are as follows:
1. Acceptor arm
- It is the site of attachment of amino acids.
- There is presence of 5’-CCA-3’ sequence at extreme 3’ end of the molecules.
- The 5’ end consists of guanine or cytosine.
2. TΨc loop
- Two unusual bases ribothymidine (T) and pseudouridine (ΨU) along with cytosine are found in this loop.
- U is found within the sequence 5’-TΨUCG-3’.
3. D loop
- Dihydrouridine is the unusual base in this loop.
- The synthetase which recognizes the amino acid activating enzyme is located on the part of D loop and apart of acceptor stem on 5’.
4. Anti-codon loop
- It has 3 nucleotides long decoding element for recognizing codon in mRNA.
- Its 3’ end has purine and 5’ end has uracil.
5. Variable loop
- It has 3 to 21 bases and is present between anticodon and ΨU loop.
- It is not present in all t-RNAs.
L- shaped: Three-Dimensional Structure
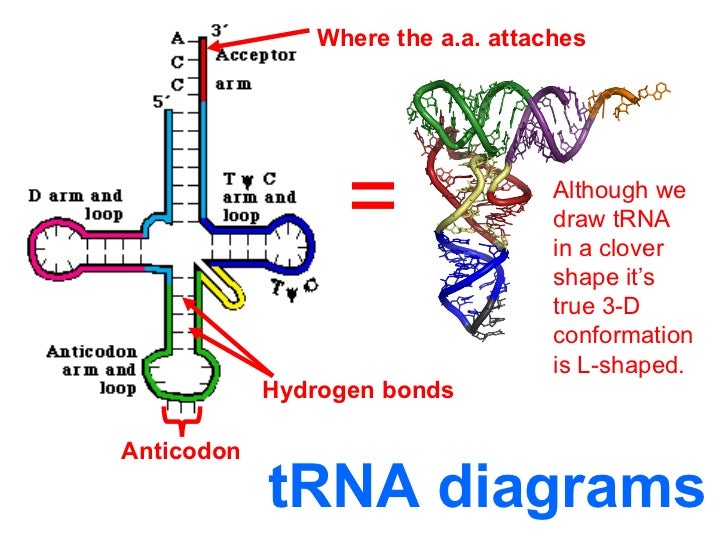
- X-ray crystallography revealed the L-shaped tertiary structure of t-RNA which has acceptor arm at one end and anticodon loop 70A away at other end.
- It is stabilized by 3 interactions. They are:
- H bonds (non-watson Crick model).
- Between bases and sugar phosphate
- Additional base stacking gained from formation of 2 extended region of base pairing.
References:
i) https://www.nature.com/scitable/definition/trna-transfer-rna-256/
ii) https://study.com/learn/lesson/trna-structure-function-synthesis.html

